aggregating for for data frames
Usage
daggregate(
data,
y = NULL,
x = NULL,
subset,
...,
fun = "summary",
regex = mets.options()$regex,
missing = FALSE,
remove.empty = FALSE,
matrix = FALSE,
silent = FALSE,
na.action = na.pass,
convert = NULL
)Arguments
- data
data.frame
- y
name of variable, or formula, or names of variables on data frame.
- x
name of variable, or formula, or names of variables on data frame.
- subset
subset expression
- ...
additional arguments to lower level functions
- fun
function defining aggregation
- regex
interpret x,y as regular expressions
- missing
Missing used in groups (x)
- remove.empty
remove empty groups from output
- matrix
if TRUE a matrix is returned instead of an array
- silent
suppress messages
- na.action
How model.frame deals with 'NA's
- convert
if TRUE try to coerce result into matrix. Can also be a user-defined function
Examples
data("sTRACE")
daggregate(iris, "^.e.al", x="Species", fun=cor, regex=TRUE)
#> Species: setosa
#> Sepal.Length Sepal.Width Petal.Length Petal.Width
#> Sepal.Length 1.0000000 0.7425467 0.2671758 0.2780984
#> Sepal.Width 0.7425467 1.0000000 0.1777000 0.2327520
#> Petal.Length 0.2671758 0.1777000 1.0000000 0.3316300
#> Petal.Width 0.2780984 0.2327520 0.3316300 1.0000000
#> ------------------------------------------------------------
#> Species: versicolor
#> Sepal.Length Sepal.Width Petal.Length Petal.Width
#> Sepal.Length 1.0000000 0.5259107 0.7540490 0.5464611
#> Sepal.Width 0.5259107 1.0000000 0.5605221 0.6639987
#> Petal.Length 0.7540490 0.5605221 1.0000000 0.7866681
#> Petal.Width 0.5464611 0.6639987 0.7866681 1.0000000
#> ------------------------------------------------------------
#> Species: virginica
#> Sepal.Length Sepal.Width Petal.Length Petal.Width
#> Sepal.Length 1.0000000 0.4572278 0.8642247 0.2811077
#> Sepal.Width 0.4572278 1.0000000 0.4010446 0.5377280
#> Petal.Length 0.8642247 0.4010446 1.0000000 0.3221082
#> Petal.Width 0.2811077 0.5377280 0.3221082 1.0000000
daggregate(iris, Sepal.Length+Petal.Length ~Species, fun=summary)
#> Species: setosa
#> Sepal.Length Petal.Length
#> Min. :4.300 Min. :1.000
#> 1st Qu.:4.800 1st Qu.:1.400
#> Median :5.000 Median :1.500
#> Mean :5.006 Mean :1.462
#> 3rd Qu.:5.200 3rd Qu.:1.575
#> Max. :5.800 Max. :1.900
#> ------------------------------------------------------------
#> Species: versicolor
#> Sepal.Length Petal.Length
#> Min. :4.900 Min. :3.00
#> 1st Qu.:5.600 1st Qu.:4.00
#> Median :5.900 Median :4.35
#> Mean :5.936 Mean :4.26
#> 3rd Qu.:6.300 3rd Qu.:4.60
#> Max. :7.000 Max. :5.10
#> ------------------------------------------------------------
#> Species: virginica
#> Sepal.Length Petal.Length
#> Min. :4.900 Min. :4.500
#> 1st Qu.:6.225 1st Qu.:5.100
#> Median :6.500 Median :5.550
#> Mean :6.588 Mean :5.552
#> 3rd Qu.:6.900 3rd Qu.:5.875
#> Max. :7.900 Max. :6.900
daggregate(iris, log(Sepal.Length)+I(Petal.Length>1.5) ~ Species,
fun=summary)
#> Species: setosa
#> log(Sepal.Length) I(Petal.Length > 1.5)
#> Min. :1.459 Mode :logical
#> 1st Qu.:1.569 FALSE:37
#> Median :1.609 TRUE :13
#> Mean :1.608
#> 3rd Qu.:1.649
#> Max. :1.758
#> ------------------------------------------------------------
#> Species: versicolor
#> log(Sepal.Length) I(Petal.Length > 1.5)
#> Min. :1.589 Mode:logical
#> 1st Qu.:1.723 TRUE:50
#> Median :1.775
#> Mean :1.777
#> 3rd Qu.:1.841
#> Max. :1.946
#> ------------------------------------------------------------
#> Species: virginica
#> log(Sepal.Length) I(Petal.Length > 1.5)
#> Min. :1.589 Mode:logical
#> 1st Qu.:1.829 TRUE:50
#> Median :1.872
#> Mean :1.881
#> 3rd Qu.:1.932
#> Max. :2.067
daggregate(iris, "*Length*", x="Species", fun=head)
#> Species: setosa
#> Sepal.Length Petal.Length
#> 1 5.1 1.4
#> 2 4.9 1.4
#> 3 4.7 1.3
#> 4 4.6 1.5
#> 5 5.0 1.4
#> 6 5.4 1.7
#> ------------------------------------------------------------
#> Species: versicolor
#> Sepal.Length Petal.Length
#> 51 7.0 4.7
#> 52 6.4 4.5
#> 53 6.9 4.9
#> 54 5.5 4.0
#> 55 6.5 4.6
#> 56 5.7 4.5
#> ------------------------------------------------------------
#> Species: virginica
#> Sepal.Length Petal.Length
#> 101 6.3 6.0
#> 102 5.8 5.1
#> 103 7.1 5.9
#> 104 6.3 5.6
#> 105 6.5 5.8
#> 106 7.6 6.6
daggregate(iris, "^.e.al", x="Species", fun=tail, regex=TRUE)
#> Species: setosa
#> Sepal.Length Sepal.Width Petal.Length Petal.Width
#> 45 5.1 3.8 1.9 0.4
#> 46 4.8 3.0 1.4 0.3
#> 47 5.1 3.8 1.6 0.2
#> 48 4.6 3.2 1.4 0.2
#> 49 5.3 3.7 1.5 0.2
#> 50 5.0 3.3 1.4 0.2
#> ------------------------------------------------------------
#> Species: versicolor
#> Sepal.Length Sepal.Width Petal.Length Petal.Width
#> 95 5.6 2.7 4.2 1.3
#> 96 5.7 3.0 4.2 1.2
#> 97 5.7 2.9 4.2 1.3
#> 98 6.2 2.9 4.3 1.3
#> 99 5.1 2.5 3.0 1.1
#> 100 5.7 2.8 4.1 1.3
#> ------------------------------------------------------------
#> Species: virginica
#> Sepal.Length Sepal.Width Petal.Length Petal.Width
#> 145 6.7 3.3 5.7 2.5
#> 146 6.7 3.0 5.2 2.3
#> 147 6.3 2.5 5.0 1.9
#> 148 6.5 3.0 5.2 2.0
#> 149 6.2 3.4 5.4 2.3
#> 150 5.9 3.0 5.1 1.8
daggregate(sTRACE, status~ diabetes, fun=table)
#> diabetes: 0
#> status
#> 0 7 9
#> 220 5 228
#> ------------------------------------------------------------
#> diabetes: 1
#> status
#> 0 9
#> 16 31
daggregate(sTRACE, status~ diabetes+sex, fun=table)
#> diabetes: 0
#> sex: 0
#> status
#> 0 9
#> 63 80
#> ------------------------------------------------------------
#> diabetes: 1
#> sex: 0
#> status
#> 0 9
#> 6 13
#> ------------------------------------------------------------
#> diabetes: 0
#> sex: 1
#> status
#> 0 7 9
#> 157 5 148
#> ------------------------------------------------------------
#> diabetes: 1
#> sex: 1
#> status
#> 0 9
#> 10 18
daggregate(sTRACE, status + diabetes+sex ~ vf+I(wmi>1.4), fun=table)
#> vf: 0
#> I(wmi > 1.4): FALSE
#> , , sex = 0
#>
#> diabetes
#> status 0 1
#> 0 21 3
#> 7 0 0
#> 9 39 8
#>
#> , , sex = 1
#>
#> diabetes
#> status 0 1
#> 0 48 6
#> 7 1 0
#> 9 94 14
#>
#> ------------------------------------------------------------
#> vf: 1
#> I(wmi > 1.4): FALSE
#> , , sex = 0
#>
#> diabetes
#> status 0 1
#> 0 2 0
#> 9 5 1
#>
#> , , sex = 1
#>
#> diabetes
#> status 0 1
#> 0 4 0
#> 9 8 0
#>
#> ------------------------------------------------------------
#> vf: 0
#> I(wmi > 1.4): TRUE
#> , , sex = 0
#>
#> diabetes
#> status 0 1
#> 0 38 3
#> 7 0 0
#> 9 34 4
#>
#> , , sex = 1
#>
#> diabetes
#> status 0 1
#> 0 102 4
#> 7 4 0
#> 9 44 4
#>
#> ------------------------------------------------------------
#> vf: 1
#> I(wmi > 1.4): TRUE
#> , , sex = 0
#>
#> diabetes
#> status 0
#> 0 2
#> 9 2
#>
#> , , sex = 1
#>
#> diabetes
#> status 0
#> 0 3
#> 9 2
#>
daggregate(iris, "^.e.al", x="Species",regex=TRUE)
#> Species: setosa
#> Sepal.Length Sepal.Width Petal.Length Petal.Width
#> Min. :4.300 Min. :2.300 Min. :1.000 Min. :0.100
#> 1st Qu.:4.800 1st Qu.:3.200 1st Qu.:1.400 1st Qu.:0.200
#> Median :5.000 Median :3.400 Median :1.500 Median :0.200
#> Mean :5.006 Mean :3.428 Mean :1.462 Mean :0.246
#> 3rd Qu.:5.200 3rd Qu.:3.675 3rd Qu.:1.575 3rd Qu.:0.300
#> Max. :5.800 Max. :4.400 Max. :1.900 Max. :0.600
#> ------------------------------------------------------------
#> Species: versicolor
#> Sepal.Length Sepal.Width Petal.Length Petal.Width
#> Min. :4.900 Min. :2.000 Min. :3.00 Min. :1.000
#> 1st Qu.:5.600 1st Qu.:2.525 1st Qu.:4.00 1st Qu.:1.200
#> Median :5.900 Median :2.800 Median :4.35 Median :1.300
#> Mean :5.936 Mean :2.770 Mean :4.26 Mean :1.326
#> 3rd Qu.:6.300 3rd Qu.:3.000 3rd Qu.:4.60 3rd Qu.:1.500
#> Max. :7.000 Max. :3.400 Max. :5.10 Max. :1.800
#> ------------------------------------------------------------
#> Species: virginica
#> Sepal.Length Sepal.Width Petal.Length Petal.Width
#> Min. :4.900 Min. :2.200 Min. :4.500 Min. :1.400
#> 1st Qu.:6.225 1st Qu.:2.800 1st Qu.:5.100 1st Qu.:1.800
#> Median :6.500 Median :3.000 Median :5.550 Median :2.000
#> Mean :6.588 Mean :2.974 Mean :5.552 Mean :2.026
#> 3rd Qu.:6.900 3rd Qu.:3.175 3rd Qu.:5.875 3rd Qu.:2.300
#> Max. :7.900 Max. :3.800 Max. :6.900 Max. :2.500
dlist(iris,Petal.Length+Sepal.Length ~ Species |Petal.Length>1.3 & Sepal.Length>5,
n=list(1:3,-(3:1)))
#> Species: setosa
#> Petal.Length Sepal.Length
#> 1 1.4 5.1
#> 6 1.7 5.4
#> 11 1.5 5.4
#> ---
#> 45 1.9 5.1
#> 47 1.6 5.1
#> 49 1.5 5.3
#> ------------------------------------------------------------
#> Species: versicolor
#> Petal.Length Sepal.Length
#> 51 4.7 7.0
#> 52 4.5 6.4
#> 53 4.9 6.9
#> ---
#> 98 4.3 6.2
#> 99 3.0 5.1
#> 100 4.1 5.7
#> ------------------------------------------------------------
#> Species: virginica
#> Petal.Length Sepal.Length
#> 101 6.0 6.3
#> 102 5.1 5.8
#> 103 5.9 7.1
#> ---
#> 148 5.2 6.5
#> 149 5.4 6.2
#> 150 5.1 5.9
daggregate(iris, I(Sepal.Length>7)~Species | I(Petal.Length>1.5))
#> Species: setosa
#> I(Sepal.Length > 7)
#> Mode :logical
#> FALSE:13
#> ------------------------------------------------------------
#> Species: versicolor
#> I(Sepal.Length > 7)
#> Mode :logical
#> FALSE:50
#> ------------------------------------------------------------
#> Species: virginica
#> I(Sepal.Length > 7)
#> Mode :logical
#> FALSE:38
#> TRUE :12
daggregate(iris, I(Sepal.Length>7)~Species | I(Petal.Length>1.5),
fun=table)
#> Species: setosa
#> I(Sepal.Length > 7)
#> FALSE
#> 13
#> ------------------------------------------------------------
#> Species: versicolor
#> I(Sepal.Length > 7)
#> FALSE
#> 50
#> ------------------------------------------------------------
#> Species: virginica
#> I(Sepal.Length > 7)
#> FALSE TRUE
#> 38 12
dsum(iris, .~Species, matrix=TRUE, missing=TRUE)
#> Species Sepal.Length Sepal.Width Petal.Length Petal.Width
#> 1 setosa 250.3 171.4 73.1 12.3
#> 2 versicolor 296.8 138.5 213.0 66.3
#> 3 virginica 329.4 148.7 277.6 101.3
par(mfrow=c(1,2))
data(iris)
drename(iris) <- ~.
daggregate(iris,'sepal*'~species|species!="virginica",fun=plot)
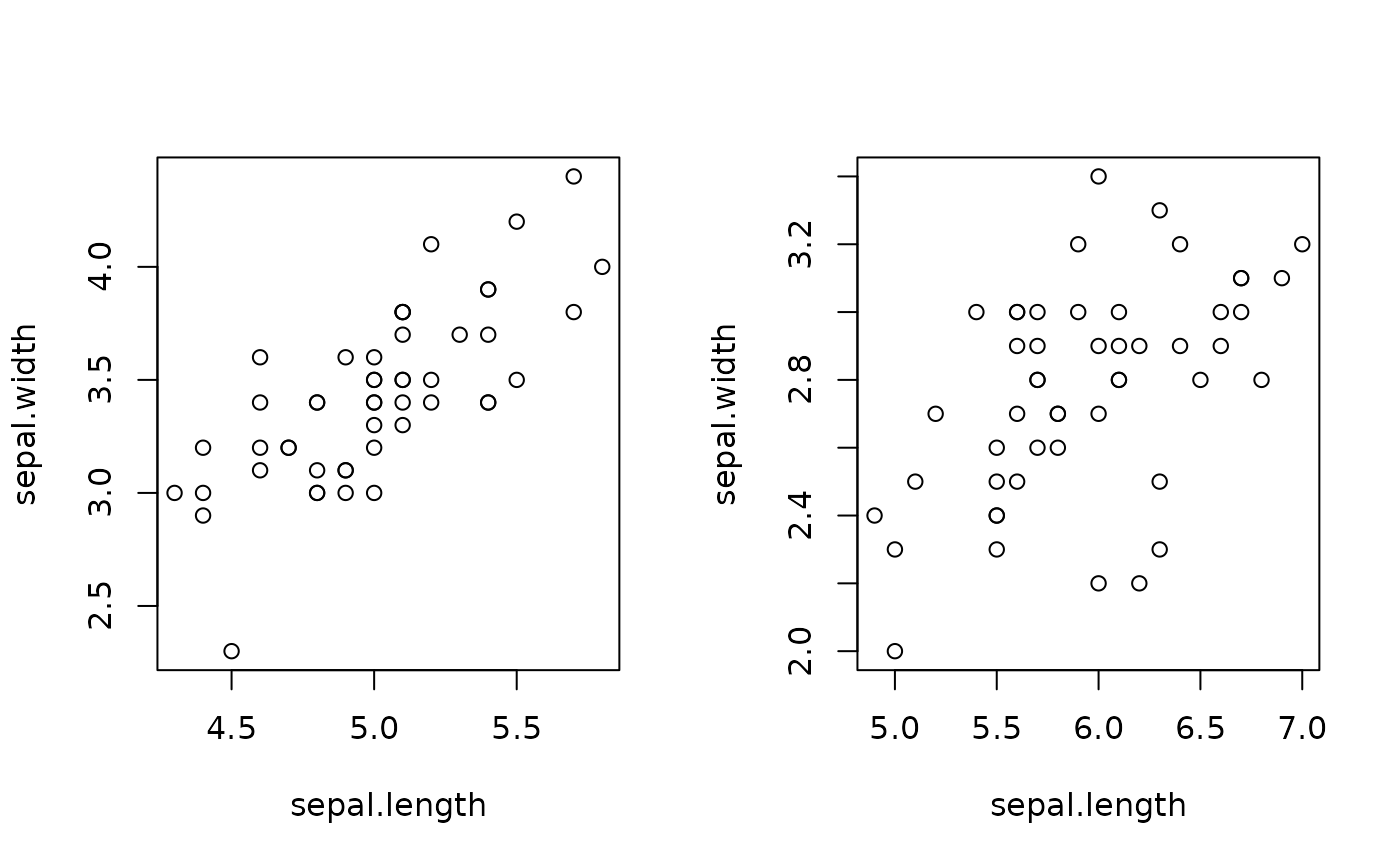 #> species: setosa
#> NULL
#> ------------------------------------------------------------
#> species: versicolor
#> NULL
#> ------------------------------------------------------------
#> species: virginica
#> NULL
daggregate(iris,'sepal*'~I(as.numeric(species))|I(as.numeric(species))!=1,fun=summary)
#> I(as.numeric(species)): 2
#> sepal.length sepal.width
#> Min. :4.900 Min. :2.000
#> 1st Qu.:5.600 1st Qu.:2.525
#> Median :5.900 Median :2.800
#> Mean :5.936 Mean :2.770
#> 3rd Qu.:6.300 3rd Qu.:3.000
#> Max. :7.000 Max. :3.400
#> ------------------------------------------------------------
#> I(as.numeric(species)): 3
#> sepal.length sepal.width
#> Min. :4.900 Min. :2.200
#> 1st Qu.:6.225 1st Qu.:2.800
#> Median :6.500 Median :3.000
#> Mean :6.588 Mean :2.974
#> 3rd Qu.:6.900 3rd Qu.:3.175
#> Max. :7.900 Max. :3.800
dnumeric(iris) <- ~species
daggregate(iris,'sepal*'~species.n|species.n!=1,fun=summary)
#> species.n: 2
#> sepal.length sepal.width
#> Min. :4.900 Min. :2.000
#> 1st Qu.:5.600 1st Qu.:2.525
#> Median :5.900 Median :2.800
#> Mean :5.936 Mean :2.770
#> 3rd Qu.:6.300 3rd Qu.:3.000
#> Max. :7.000 Max. :3.400
#> ------------------------------------------------------------
#> species.n: 3
#> sepal.length sepal.width
#> Min. :4.900 Min. :2.200
#> 1st Qu.:6.225 1st Qu.:2.800
#> Median :6.500 Median :3.000
#> Mean :6.588 Mean :2.974
#> 3rd Qu.:6.900 3rd Qu.:3.175
#> Max. :7.900 Max. :3.800
#> species: setosa
#> NULL
#> ------------------------------------------------------------
#> species: versicolor
#> NULL
#> ------------------------------------------------------------
#> species: virginica
#> NULL
daggregate(iris,'sepal*'~I(as.numeric(species))|I(as.numeric(species))!=1,fun=summary)
#> I(as.numeric(species)): 2
#> sepal.length sepal.width
#> Min. :4.900 Min. :2.000
#> 1st Qu.:5.600 1st Qu.:2.525
#> Median :5.900 Median :2.800
#> Mean :5.936 Mean :2.770
#> 3rd Qu.:6.300 3rd Qu.:3.000
#> Max. :7.000 Max. :3.400
#> ------------------------------------------------------------
#> I(as.numeric(species)): 3
#> sepal.length sepal.width
#> Min. :4.900 Min. :2.200
#> 1st Qu.:6.225 1st Qu.:2.800
#> Median :6.500 Median :3.000
#> Mean :6.588 Mean :2.974
#> 3rd Qu.:6.900 3rd Qu.:3.175
#> Max. :7.900 Max. :3.800
dnumeric(iris) <- ~species
daggregate(iris,'sepal*'~species.n|species.n!=1,fun=summary)
#> species.n: 2
#> sepal.length sepal.width
#> Min. :4.900 Min. :2.000
#> 1st Qu.:5.600 1st Qu.:2.525
#> Median :5.900 Median :2.800
#> Mean :5.936 Mean :2.770
#> 3rd Qu.:6.300 3rd Qu.:3.000
#> Max. :7.000 Max. :3.400
#> ------------------------------------------------------------
#> species.n: 3
#> sepal.length sepal.width
#> Min. :4.900 Min. :2.200
#> 1st Qu.:6.225 1st Qu.:2.800
#> Median :6.500 Median :3.000
#> Mean :6.588 Mean :2.974
#> 3rd Qu.:6.900 3rd Qu.:3.175
#> Max. :7.900 Max. :3.800