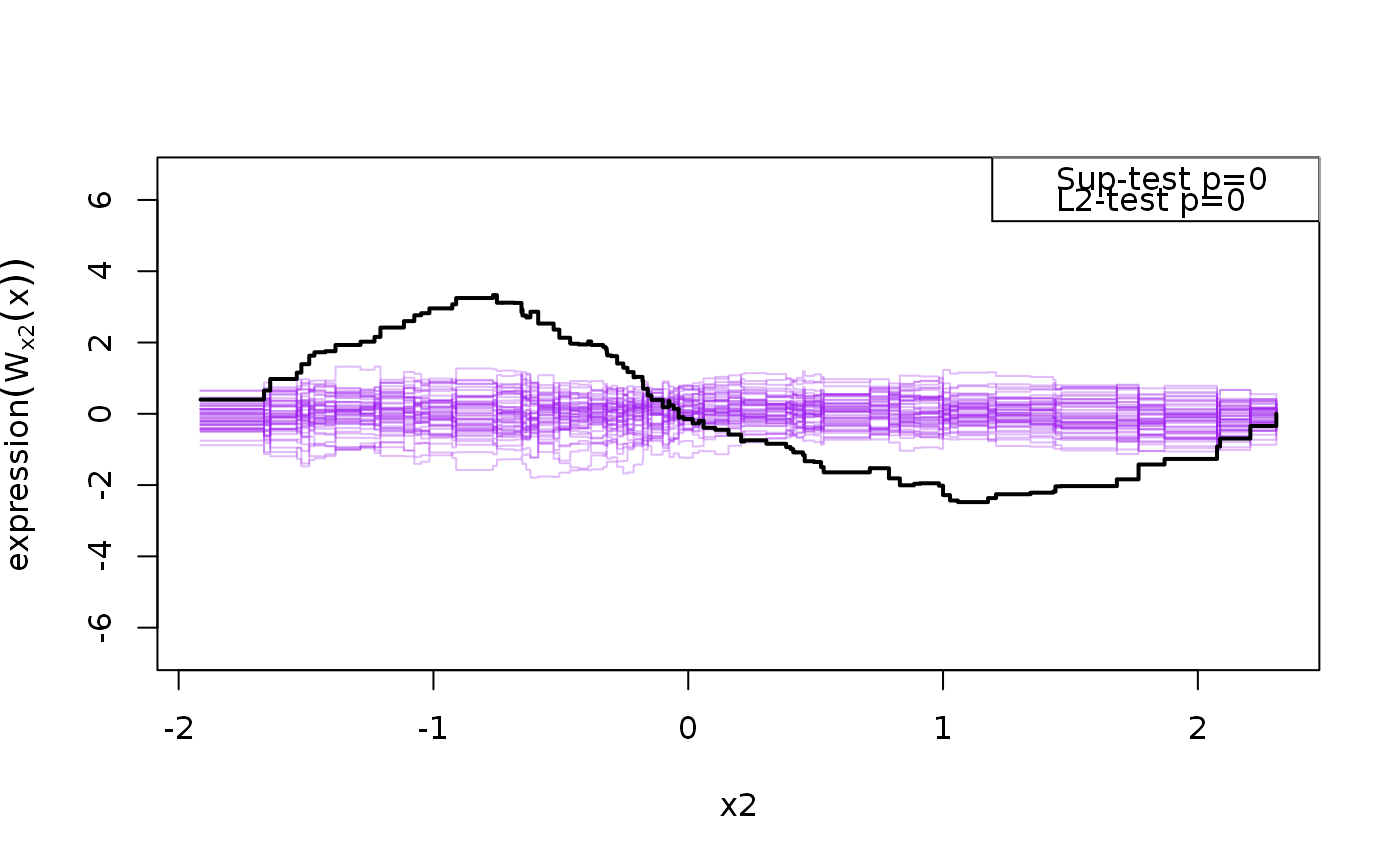
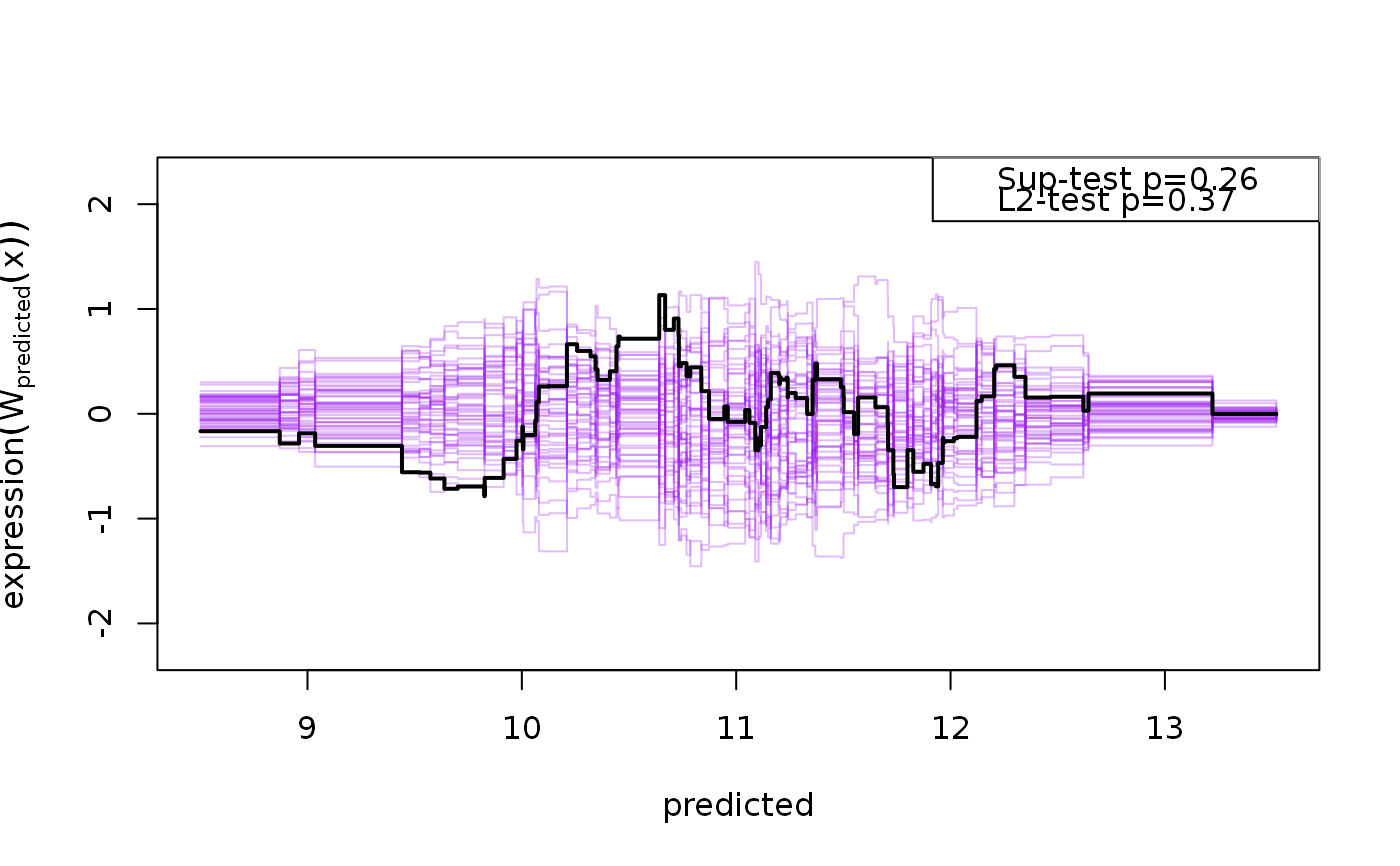
Calculates GoF statistics based on cumulative residual processes
cumres.glm.RdGiven the generalized linear models model $$g(E(Y_i|X_{i1},...,X_{ik})) =
\sum_{i=1}^k \beta_jX_{ij}$$ the cumres-function calculates the the
observed cumulative sum of residual process, cumulating the residuals,
\(e_i\), by the jth covariate: $$W_j(t) = n^{-1/2}\sum_{i=1}^n
1_{\{X_{ij}<t\}}e_i$$ and Sup and L2 test
statistics are calculated via simulation from the asymptotic distribution of
the cumulative residual process under the null (Lin et al., 2002).
# S3 method for glm cumres( model, variable = c("predicted", colnames(model.matrix(model))), data = data.frame(model.matrix(model)), R = 1000, b = 0, plots = min(R, 50), ... )
Arguments
| model | Model object ( |
|---|---|
| variable | List of variable to order the residuals after |
| data | data.frame used to fit model (complete cases) |
| R | Number of samples used in simulation |
| b | Moving average bandwidth (0 corresponds to infinity = standard cumulated residuals) |
| plots | Number of realizations to save for use in the plot-routine |
| ... | additional arguments |
Value
Returns an object of class 'cumres'.
Note
Currently linear (normal), logistic and poisson regression models with canonical links are supported.
References
D.Y. Lin and L.J. Wei and Z. Ying (2002) Model-Checking Techniques Based on Cumulative Residuals. Biometrics, Volume 58, pp 1-12.
John Q. Su and L.J. Wei (1991) A lack-of-fit test for the mean function in a generalized linear model. Journal. Amer. Statist. Assoc., Volume 86, Number 414, pp 420-426.
See also
cox.aalen in the timereg-package for
similar GoF-methods for survival-data.
Author
Klaus K. Holst
Examples
sim1 <- function(n=100, f=function(x1,x2) {10+x1+x2^2}, sd=1, seed=1) { if (!is.null(seed)) set.seed(seed) x1 <- rnorm(n); x2 <- rnorm(n) X <- cbind(1,x1,x2) y <- f(x1,x2) + rnorm(n,sd=sd) d <- data.frame(y,x1,x2) return(d) } d <- sim1(100); l <- lm(y ~ x1 + x2,d) system.time(g <- cumres(l, R=100, plots=50))#> user system elapsed #> 0.023 0.001 0.023g#> #> p-value(Sup) p-value(L2) #> predicted 0.26 0.37 #> x1 0.47 0.42 #> x2 0.00 0.00 #> #> Based on 100 realizations.plot(g)#> #> p-value(Sup) p-value(L2) #> predicted 0.35 0.35 #> #> Based on 100 realizations.#> #> p-value(Sup) p-value(L2) #> predicted 0.21 0.26 #> #> Based on 100 realizations.